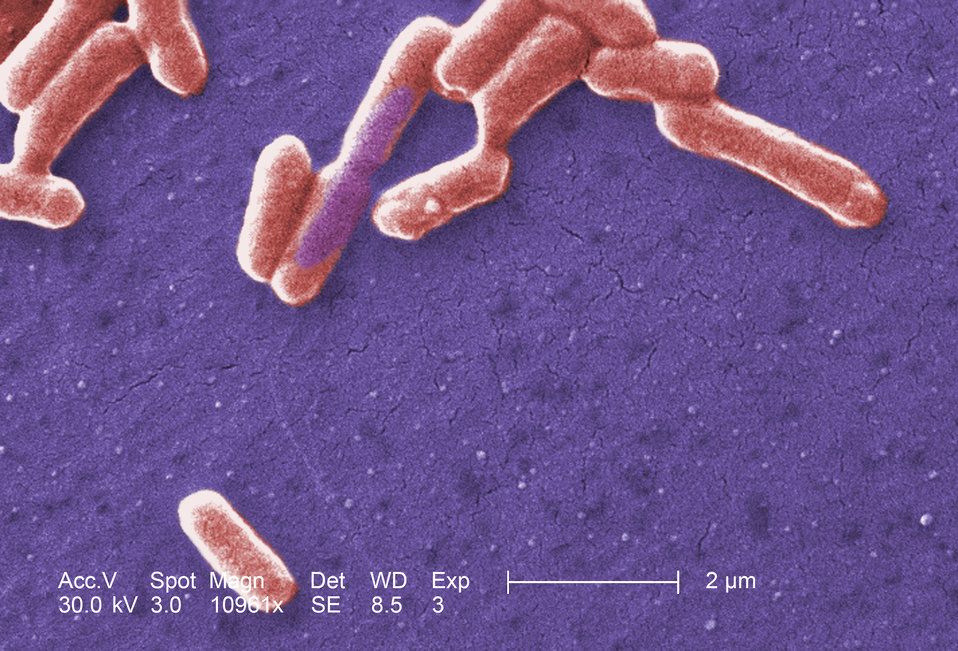
The small bacteria Escherichia coli is a well known and commonly used model organism in science. As such, it makes an excellent organism for us to use to show some of the problems with evolutionary genetics. In order to understand this article, you will either need knowledge of genome evolution or have read our previous article on evolutionary novelty.
The E. coli genome is not a large genome. This makes sense given it is a very small organism. The genome is only 4.6 million base pairs long, with not quite 4300 protein-coding genes (Blattner et al, 1997). Now the evolutionists are undoubtedly ignoring the regulatory elements in the E. coli genome, but we’re going to play by their rules here and give them every benefit of the doubt like we always do.
Since we know that the genome can be read in both directions and many genes can be expressed multiple ways (Buchanan et al, 2009), evolution has a lot of complexity to account for, even in something as small as E. coli. Given that a study of E. coli revealed that only 1 in 150 mutations are beneficial (Gordo et al, 2007) and neofunctionalization requires beneficial mutations, it is almost certainly uncommon. Again, we are giving the evolutionists the benefit of the doubt here and assuming that these are beneficial mutations, not simply broken information that happens to be situationally beneficial. Nonfunctionalization is the most likely outcome of a duplicated gene given the aforementioned data if mutations are allowed to accumulate.
However, if mutation rate is low enough, subfunctionalization and gene conservation can come into play. The mutation rate in E. coli is 8.9 x e-11 per nucleotide, per generation (Schneider et al, 2011) The generation time of E. coli varies depending on conditions so we will take the approach best suited to evolution, thirty minutes (Planck and Harvey, 1979). This is under ideal conditions constantly, since E. coli evolved. It is also a wildly unrealistic expectation that conditions have been ideal since time immemorial, but, again, we are giving evolution the benefit of the doubt. Using simple math, that means E. coli undergoes 17, 520 generations per year at a maximum. The E. coli genome is 4,639,221 base pairs long (Blattner et al, 1997). Double that to account for nucleotides on each end of the base pair yields 9,278,442 nucleotides in the genome.
This is where the math gets interesting.
0.000000000089 mutation rate/1nucleotide x 9,278,442 nucleotides/1genome =8.27x e-4 mutations rate per genome per generation
8.27x e-4 mutations rate per genome/ 1 generation x 17,520 generations/ 1 year = 14.5 new mutations per lineage per year.
Since we know only 1/150 mutations are beneficial in E. coli and we have now deduced the number of new mutations in an E.coli lineage that occur per year, we can determine how many years it will take to produce one beneficial mutation on average.
150 mutations/14.5 mutations yearly= 10.3 years for one beneficial mutation to occur.
Since we know the number of generations E. coli conservatively undergoes per year, we can determine how many generations of E. coli must pass before that beneficial mutation occurs.
17, 520 generations per year x10.3 years= 180,456 generations for one beneficial mutation to occur. It would then need to become fixed in the population, an outcome that is far from guaranteed.
Just by working through the numbers, I think we can demonstrate how rare neofunctionalization must be and how common nonfunctionalization must be. Remember too, the evolutionists don’t just need one or two beneficial mutations once in a while. They need thousands, perhaps even hundreds of thousands to generate all the new information for new phenotypes. In this model, we have given them every possible benefit of the doubt. To get just one beneficial mutation requires over one hundred and eighty thousand generations. Given that most traits are polygenetic, this is simply not fast enough. Evolution cannot explain the origin of new traits genetically. The math simply does not allow it.
References:
Blattner FR, Plunkett G, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF. 1997. The Complete Genome Sequence of Escherichia coli K-12. Science (80- ). 277(5331):1453-1462.
Buchanan AV, Sholtis S, Richtsmeier J, Weiss KM. 2009. What are genes “for” or where are traits “from”? What is the question?. Bioessays. 31(2):198-208.
Gordo I, Perfeito L, Fernandes L, Mota C. 2007. Adaptive mutations in bacteria: high rate and small effects. Science. 317(5839):813-815.
Plank LD, Harvey JD. 1979. Generation time statistics of Escherichia coli B measured by synchronous culture techniques. J of Microbiology. 115(1): 69-77.
Schneider D, Wielgoss S, Barrick JE, Tenaillon O, Cruveiller S, Chance-Woon-Ming B, Médigue C, Lenski RE. 2011. Mutation rate inferred from synonymous substitutions in a long term evolution experiment with Escherichia coli. G3 (Bethesda). 1(3):183-186.