In this ongoing series, we have been walking through the McLain et al attempt to feather dinosaurs using baraminology. So far, after looking at the data, I am significantly underwhelmed. None of the graphs presented have provided clear evidence of dinosaurs having feathers. Frankly, the whole thing feels like smoke and mirrors. However, we will continue to work through their data in the hopes of finding some support for their position baraminologically. In this article, we look at figures 36 and 37 from McLain et al, 2018.
As usual, the first step was to determine bootstrapping values as these were not reported in the paper. The group we examined this week is the Paraves subset of the Lee dataset,exactly as run by McLain et al. Fourteen taxa were used, with a total of 277 characters retained for analysis at 75% relevance. Bootstrapping values were decent with quite a few in the mid 90s to 100 range. However, a few were quite poor, in the 40s and 50s. Given at least a few bootstrapping values were poor, it raises questions about how good the data is. Again I have to ask why this important step in the method was completely ignored? I understand that the paper was monstrous, but an additional paragraph or sentence explaining bootstrapping values, or why the authors elected to not report them, would not have hurt.
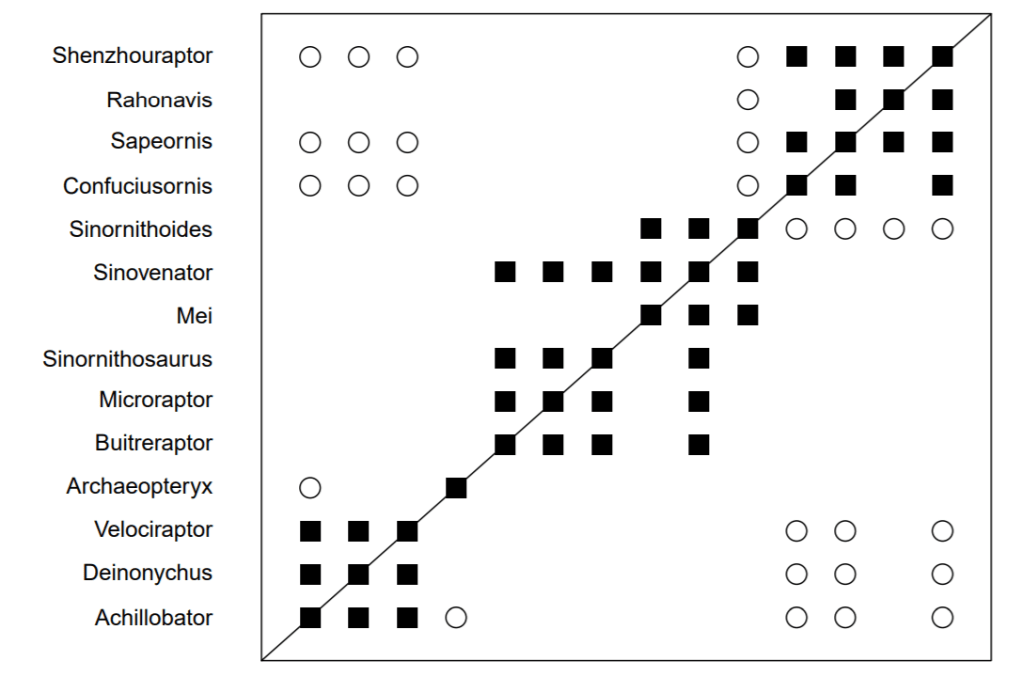
Having worked through the boostrapping values, I attempted to duplicate the graph presented in figure 36. I was shocked to discover that BDISTMDS did not replicate the graph! Using the same data, with the same number of characters, the graph I generated from BDISTMDS did not match McLain et al’s published graph. As you can see below, the generated BDIST graph shows a wall of discontinuity surrounding the topmost group. That is absent in McLain’s graph. The discontinuity wall also shows up in the BARCLAY graph. I can think of no reason for this difference unless Dr. Wood modified the algorithm of BDISTMDS between McLain publishing and the present, or McLain et al modified the algorithm. In both cases, I would like to know why this was not reported in the literature?
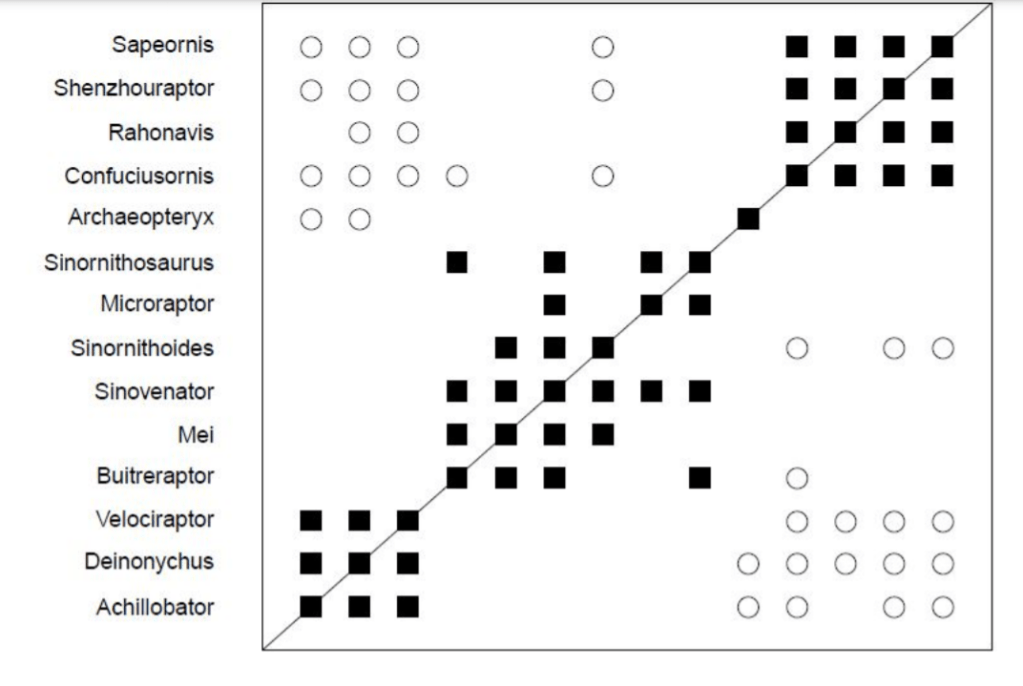



Note also about these results that, again, BARCLAY is even worse at detecting discontinuity than BDISTMDS. The bottom two blocks of taxa have been linked together where previously they were not and Archaeopteryx now links into the top block and the middle block. Effectively, this makes the whole graph join together as one baraminic group.
Out of curiosity, I reran my analysis at 85% relevance. All taxa and 154 characters were retained for analysis. The graphs were similar to the ones displayed above, with Archaeopteryx dropping out of the top block in BARCLAY, leading to at least one defined group as shown below.

Having worked through the graphs, I moved on to duplicating the MDS plots. Its worth reminding everyone still bothering to read these articles that MDS plots are purely subjective and thus meaningless, but they have been incorporated into the methodology so we will continue to use them. As you can see, the standard MDS plots look fairly similar. Both show a kind of triangular shape with three distinct groupings.


The above MDS plots seem to validate the idea that there are three distinct groups within the data. As you can see below, the results from an 85% relevance cut off are much the same. This is expected since the graphs were also much the same.


Given the above, what can we say about this subsection of the Lee data? I think this is actually somewhat sound in terms of data. The method is still a raging dumpster fire, but the data itself here works well within the method. That said, there are still significant concerns both about the inferences McLain et al draw from the baraminology, the data they used, and the method itself. We will continue to explore these deficiencies as we continue our series next week.
Do you know what’s going to happen when you die? Are you completely sure? If you aren’t, please read this or listen to this. You can know where you will spend eternity. If you have questions, please feel free to contact us, we’d love to talk to you.


